Diabetes Classification Case Studies
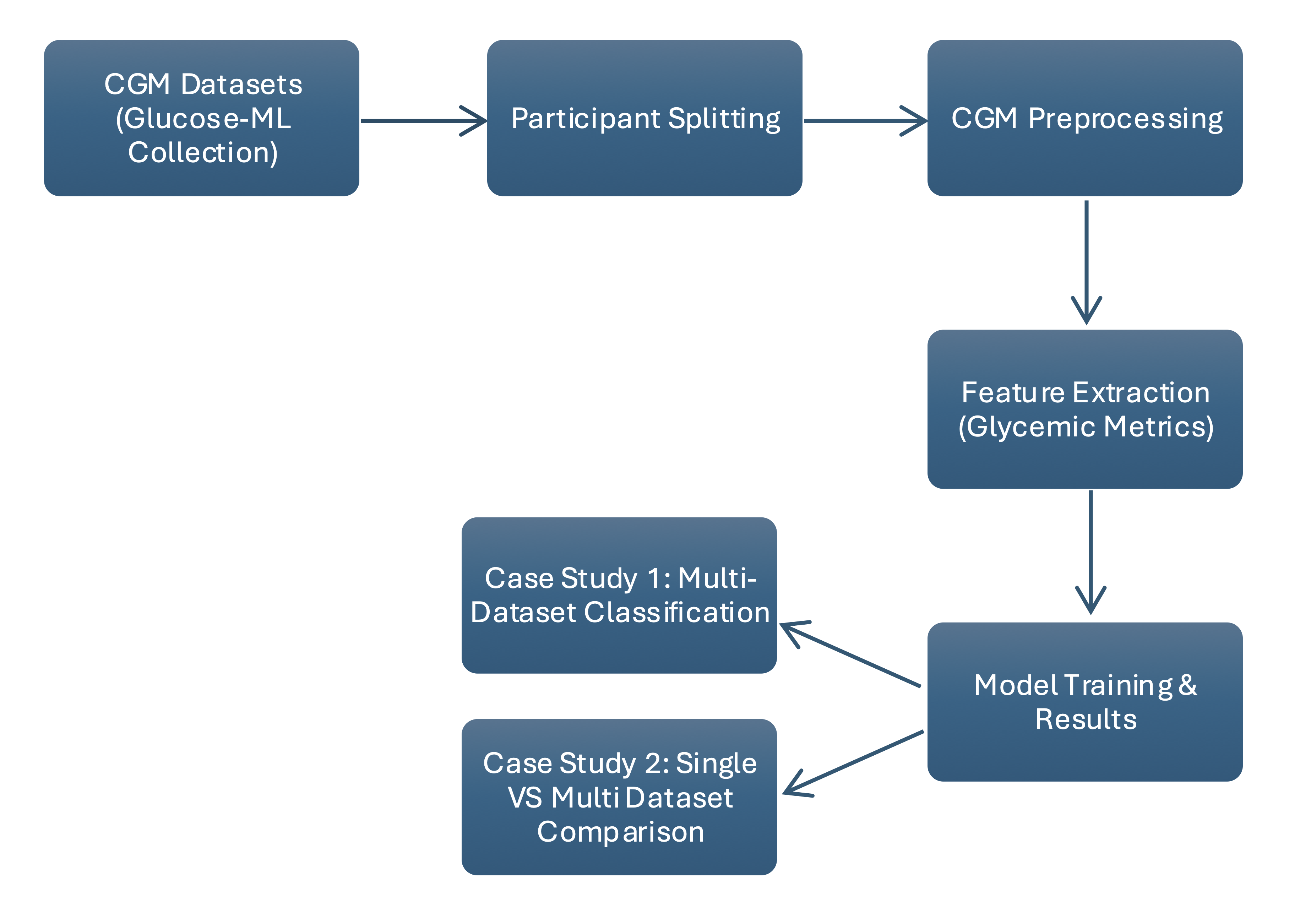
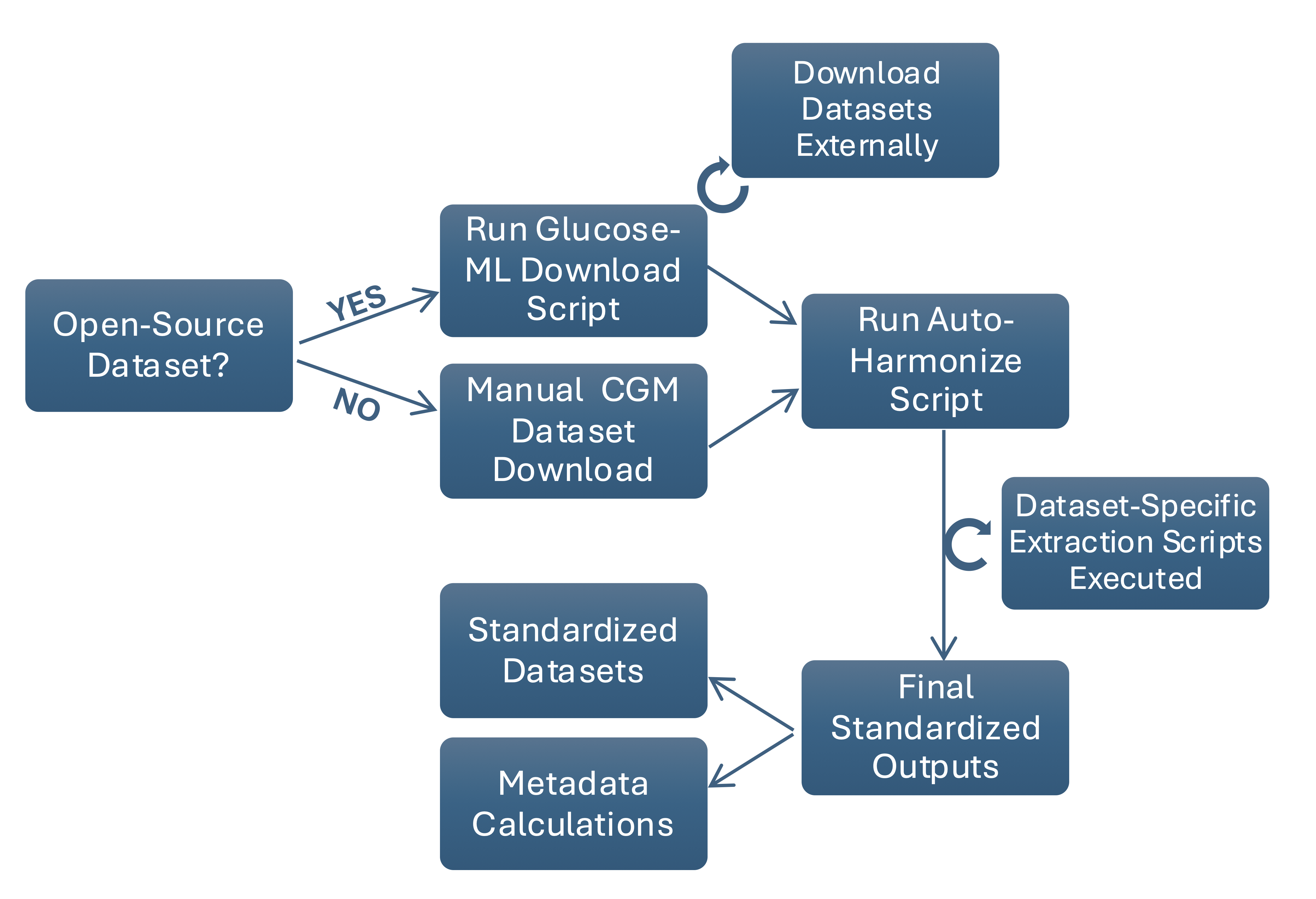
I developed two machine learning case studies using harmonized open-source CGM datasets from the Glucose-ML collection to evaluate diabetes status classification from participant-level glucose features. The first benchmark trained logistic regression, random forest, and XGBoost models across 13 datasets to classify T1D, T2D, no diabetes, and prediabetes. The second compared single-dataset versus multi-dataset training to evaluate how dataset diversity and sample size influence classification performance.
Python • Scikit-Learn • Machine Learning